Swiss Institute of Bioinformatics
Click2Drug
Directory of computer-aided Drug Design tools
Click2Drug contains a comprehensive list of computer-aided drug design (CADD) software, databases and web services. These tools are classified according to their application field, trying to cover the whole drug design pipeline. If you think that an interesting tool is missing in this list, please contact usClick on the following picture to select tools related to a given activity:
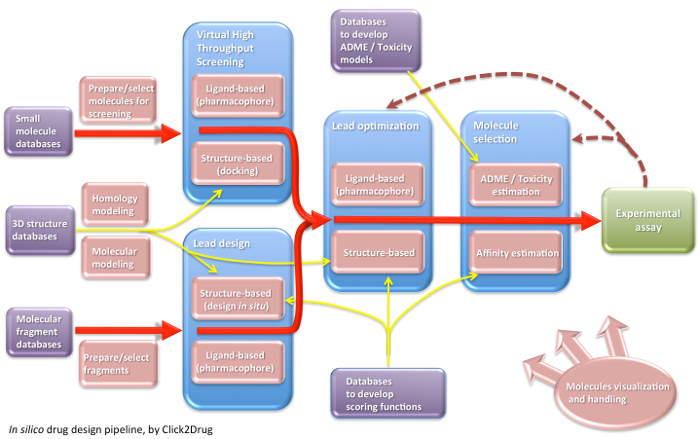
Show all links Hide all links
Databases
Chemical databases
- ChEMBL. Curated database of small molecules. Includes interactions and functional effects of small molecules binding to their macromolecular targets, and series of drug discovery databases.
Protein-ligand complexes databases
- Binding MOAD (Mother Of All Database). Subset of the Protein Data Bank (PDB), containing a collection of well resolved protein crystal structures with clearly identified biologically relevant ligands annotated with experimentally determined binding data extracted from literature. Maintained by the university of Michigan.
- PDBbind. Collection of experimentally measured binding affinity data (Kd, Ki, and IC50) exclusively for the protein-ligand complexes available in the Protein Data Bank (PDB). All of the binding affinity data compiled in this database are cited from original references.
- AffinDB. Freely accessible database of affinities for protein-ligand complexes from the PDB.
- Protein Ligand Database (PLD). Collection of protein ligand complexes extracted fom the PDB along with biomolecular data, including binding energies, Tanimoto ligand similarity scores and protein sequence similarities of protein-ligand complexes. Maintained by the University of Cambridge.
- BindingDB. Public, web-accessible database of measured binding affinities, focusing chiefly on the interactions of protein considered to be drug-targets with small, drug-like molecules.
- Ki Database. Provides information on the abilities of drugs to interact with an expanding number of molecular targets. The Ki database serves as a data warehouse for published and internally-derived Ki, or affinity, values for a large number of drugs and drug candidates at an expanding number of G-protein coupled receptors, ion channels, transporters and enzymes. Currently 55472 Ki values. Maintained by the NIMH Psychoactive Drug Screening Program.
- SCORPIO. Free online repository of protein-ligand complexes which have been structurally resolved and thermodynamically characterised.
- PDSP. Psychoactive Drug Screening Program. Provides screening of novel psychoactive compounds for pharmacological and functional activity at cloned human or rodent CNS receptors, channels, and transporters. Assays, Ki,...
- BAPPL complexes set. 161 protein-ligand complexes with experimental and estimated binding free energies calculated with the BAPPL server.
- DNA Drug complex dataset. Dataset of DNA-drug complexes consisting of 16 minimized crystal structures and 34 model-built structures, along with experimental affinities, used to validate PreDDICTA.
- Binding Database. Public, web-accessible database of measured binding affinities, focusing chiefly on the interactions of protein considered to be drug-targets with small, drug-like molecules. Maintained by the Center for Advanced Research in Biotechnology, University of Maryland Biotechnology Institute.
- Kuntz Protein Test Set. Set of 114 crystallographically determined protein-ligand structures used to test the docking program DOCK. Maintained by UCSF.